The study was led by the Science and Technology Facilities Council (STFC) Central Laser Facility (CLF) and used advanced laser imaging techniques to identify structural details of the mutated protein which help it to evade drugs that target it.
It is published today (19/03/2024) in the journal, Nature Communications and lays the groundwork for future research into more effective, long-lasting cancer therapies.
The Epidermal Growth Factor Receptor (EGFR) is a protein that sits on the surface of cells and receives molecular signals that tell the cell to grow and divide. In certain types of cancer, mutated EGFR stimulate uncontrolled growth, resulting in tumours.
Various cancer treatments block and inhibit mutant EGFR to prevent tumour formation, but these are limited as eventually cancerous cells commonly develop further EGFR mutations that are resistant to treatment.
Until now, how exactly these drug-resistant EGFR mutations drive tumour growth was not understood, hindering our ability to develop treatments that target them.
In this latest study, scientists at CLF have obtained super-resolution images of a drug-resistant EGFR mutation known to contribute to lung cancer. This was achieved using an advanced laser imaging technique developed by STFC for this purpose called Fluorophore Localisation Imaging with Photobleaching, or FLImP (pictured below).
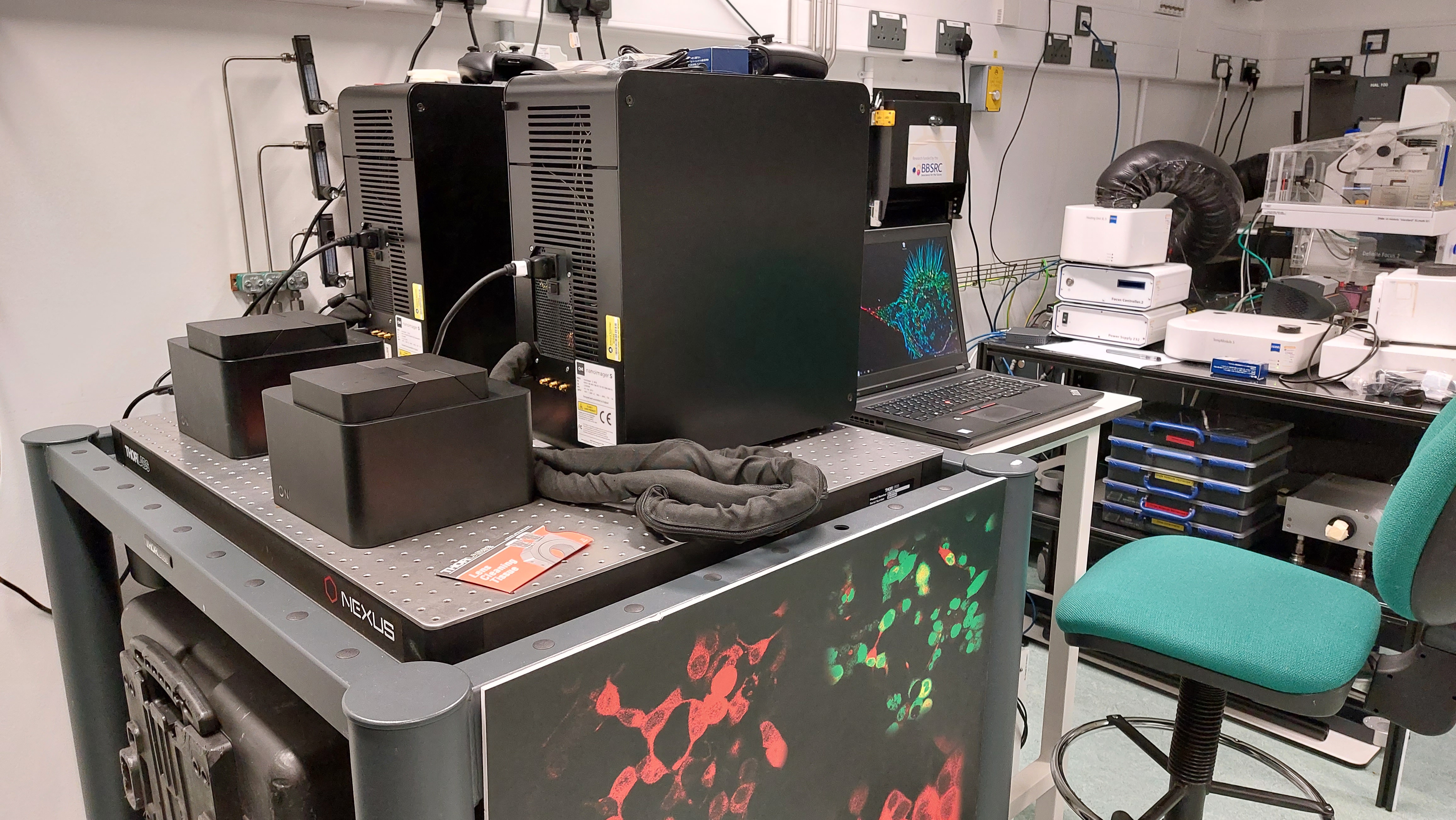

FLImP analysis revealed structural details as small as two nanometres and showed for the first time with this level of precision how the molecules in the drug-resistant EGFR mutation interact.
Additional analysis by the Biomolecular & Pharmaceutical Modelling Group at University of Geneva (UNIGE) used advanced computer simulations that combined with the FLImP analysis were able to provide atomistic details of the EGFR complexes.
From this, the team were able to compare the structural details of the mutated and healthy EGFR to identify interfaces between interacting molecules in the drug-resistant mutation critical for tumour growth.
Professor Marisa Martin-Fernandez, Leader of the CLF Octopus Group, which led the study, said:
“This finding is the culmination of years of research and technological development at CLF and our partner institutions and we’re extremely excited about its potential to inform the course of cancer research going forward. If this interface proves to be an effective therapeutic target, it could provide an entirely new approach to much needed pharmaceutical development.”
The team then introduced additional mutations to the drug-resistant EGFR in in cultured lung cells and in mice that interfered with the newly discovered interfaces.
In these experiments, one of the additional EGFR mutations was shown to block cancer growth, with mice developing no tumours, further indicating that the ability of this EGFR mutation to promote cancer depends on these interfaces.
Gilbert Fruhwirth, Leader of the Imaging Therapies and Cancer group at King's College London who validated results in live animals, said:
"This research has become possible through the combination of a variety of different imaging technologies, ranging from single molecules to whole animals, and demonstrates the power of imaging to better understand the inner workings of cancer. We are extremely pleased about this successful collaboration and look forward to develop this pharmaceutical opportunity further as part of this team.”
Researchers hope therefore that these interfaces could act as potential new targets for cancer therapies that overcome resistance acquired by EGFR mutations.
Professor Francesco Luigi Gervasio, Leader of the Biomolecular & Pharmaceutical Modelling Group at UNIGE, said:
“This breakthrough was made possible by a combination of state-of-the-art simulations and experimental techniques that can now ‘visualise’ the structure and dynamics of important cancer targets such as EGFR in unprecedented detail.”
Dr. Yiannis Galdadas at UNIGE, who performed the simulations, said:
"The simulations were able to push the effective resolution of the microscope beyond the limits of imagination. It's almost possible to ‘touch’ the mutation site and see its effect."