Their results provide promising evidence for the potential of using 2D-IR alongside ANOVA-PCA as a rapid and effective probe of DNA binding interactions.
The molecular interactions between proteins and DNA are imperative for biological function. From transcription factors that regulate gene expression levels, to the histones that package DNA into structural units called nucleosomes, these interactions are fundamental. It is for this reason that they are prime targets for drug discovery. The only problem is that current techniques used to analyse ligand-DNA binding in a sequence-specific manner, such as work with NMR or DNA footprinting experiments take longer and require more material than new 2D-IR experiments.
2D-IR or two-dimensional infrared spectroscopy is a type of multidimensional spectroscopy (the correlation of one laser excitation frequency with another) applied to the vibrational degrees of freedom of a molecule. The high time resolution of this technique makes it perfect as a tool for studying transient molecular structure and dynamics in solution. In this experiment, scientists from the University of Strathclyde were able to use 2D-IR at the CLF to gain a more information-rich view of how ligands interact with DNA compared to conventional IR techniques that have been previously used.
One of the advantages of laser systems like Lifetime with pulse repetition rates of 100 kHz is that spectral acquisition times have been reduced to a matter of seconds. This increase in the efficiency of data collection is promising for future applications of 2D-IR in a more analytical context. This might be, for example, its potential use as a tool for screening novel bioactive compounds.
The team, led by Professor Neil Hunt, measured the 2D-IR spectra of 12 double-stranded (ds) DNA oligomers in the presence and absence of binding molecule, Hoechst 33258. They did so at a series of waiting times so that they could retrieve dynamic information about ligand binding. By applying ANOVA-PCA to a 2D-IR data set for the first time, the team were able to not only demonstrate how one can retrieve the base composition of a DNA sequence but also to discriminate ligand-DNA complexes from unbound sequences.
“The ability to obtain 2D-IR spectra in a few seconds that Lifetime provides has opened up exciting new opportunities for screening-type applications. Previously, comparing ligand-biomolecule binding using a dataset containing over 2000 spectra would have been extremely time-consuming. Linking Lifetime to user-friendly data analysis tools now makes this type of experiment both possible and broadly accessible."
Professor Neil Hunt, University of Strathclyde
Excitingly, the results from this study might just pave the way for future large-scale screening tests of ligands or drugs using time-resolved 2D-IR spectroscopy. This is because the methods used are scalable and could be transferred to a range of experiments featuring multiple targets and ligands. The potential also exists to study proteins as well as DNA.
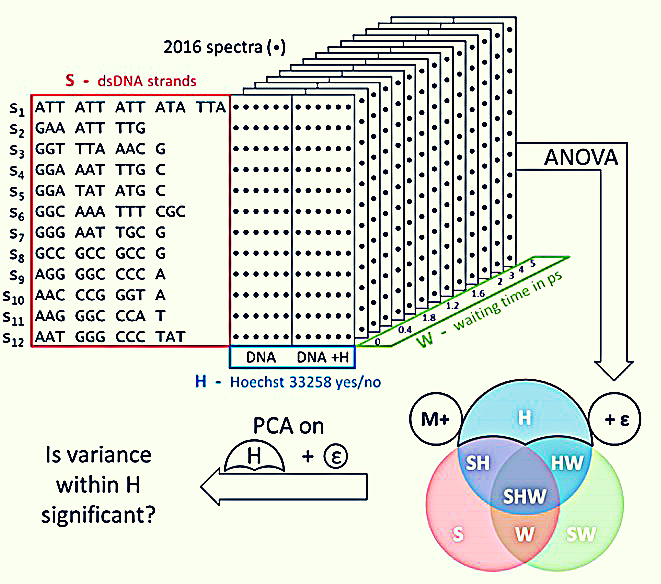
Figure 1: Experimental design and schematic representation of the ANOVA-PCA method used in this experiment. The set of 2D-IR spectra studied contains three main sources of variance (factors); the sequence of dsDNA, S, the presence of H33258, H, and the waiting time, W (Fritzsch et al, 2018)
The research was supported by STFC and the BBSRC.
The full publication is available to view in Analytical Chemistry.
For further information about the research, please contact Professor Neil Hunt (neil.hunt@strath.ac.uk)
To find out more about the capabilities of ULTRA, please follow this link.